|
6/17/2023 0 Comments Iclip experiments protocols rna ip 
For the scientific community, the revelation of the dynamicity and malleability of the partnership between RBPs and RNA allows for the exploration of new possible interaction mechanisms, networks, genes and protein regulation systems to investigate. In fact, indiscriminate binding of RNA by RBPs is a quite common phenomenon, and the assumption that stronger affinity translates into more relevant biological functions is not necessarily correct. The interactions of proteins with RNA can be highly dynamic and heavily dependent on the cellular environment, which makes the goal of defining the range of affinities and specificities quite challenging. Since one of the principal ways by which RNA exerts its function is by the formation of ribonucleoprotein complexes, every protein capable of establishing even weak and extemporary interactions with an RNA molecule may be defined as RBP. However, concomitantly to a new appreciation for RNA as key biological macromolecule acting at post-transcriptional level, there has also been a re-evaluation of what constitutes an RBP. 
Since their discovery and until recently, RNA-binding proteins (RBPs) have been identified by the presence of one or more RNA-binding domains in their sequences. The results thus obtained can furthermore validate the computational predictions, offering an all-round solution to the issue of finding the most likely RNA binding partners for a newly identified potential RBP. The RNA candidates with the highest potential can then be analysed in vitro to determine the binding strength and to precisely identify the binding sites. The large pool of potential binders selected by in-cell experiments can be enriched by in silico tools such as catRAPID, which is able to predict the RNA sequences more likely to interact with specific RBP regions with high accuracy. In this mini-review, we propose a workflow for the identification of the RNA binding partners of putative, newly identified RBPs. Numerous experimental and computational methodologies have been developed to predict, identify and verify the binding between a given RBP and potential RNA partners, but navigating across the vast ocean of data can be frustrating and misleading. However, defining the RNA sequences and structures that are selectively recognised by an RBP remains challenging, since these interactions can be transient and highly dynamic, and may be mediated by unstructured regions in the protein, as in the case of many non-canonical RBPs. The investigation of the biological relevance of non-coding RNAs has led to the identification of numerous novel RNA-binding proteins (RBPs). Sanford JR, Wang X, Mort M et al (2009) Splicing factor SFRS1 recognizes a functionally diverse landscape of RNA transcripts.Interactions between proteins and RNA are at the base of numerous cellular regulatory and functional phenomena. Yeo GW, Coufal NG, Liang TY et al (2009) An RNA code for the FOX2 splicing regulator revealed by mapping RNA-protein interactions in stem cells. Sugimoto Y, Konig J, Hussain S et al (2012) Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein-RNA interactions, Genome Biol 13:R67 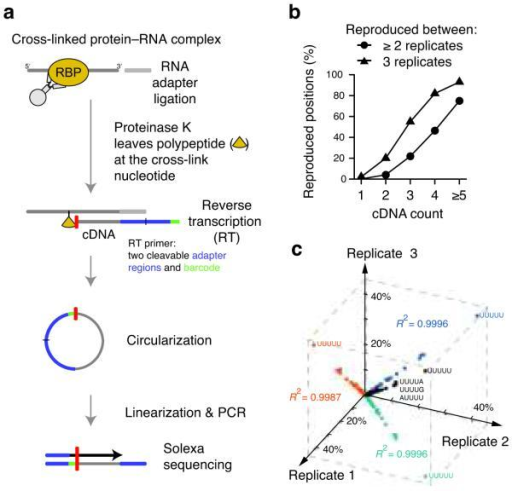
Langmead B, Trapnell C, Pop M et al (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome, Genome Biol 10:R25 Yao C, Biesinger J, Wan J et al (2012) Transcriptome-wide analyses of CstF64-RNA interactions in global regulation of mRNA alternative polyadenylation. Hafner M, Renwick N, Brown M et al (2011) RNA-ligase-dependent biases in miRNA representation in deep-sequenced small RNA cDNA libraries. Konig J, Zarnack K, Rot G et al (2010) iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Ule J, Jensen KB, Ruggiu M et al (2003) CLIP identifies Nova-regulated RNA networks in the brain. Gilbert C, Kristjuhan A, Winkler GS et al (2004) Elongator interactions with nascent mRNA revealed by RNA immunoprecipitation. Tenenbaum SA, Carson CC, Lager PJ et al (2000) Identifying mRNA subsets in messenger ribonucleoprotein complexes by using cDNA arrays. RNA 5:1071–1082īrooks SA, Rigby WF (2000) Characterization of the mRNA ligands bound by the RNA binding protein hnRNP A2 utilizing a novel in vivo technique. Trifillis P, Day N, Kiledjian M (1999) Finding the right RNA: identification of cellular mRNA substrates for RNA-binding proteins. Hieronymus H, Silver PA (2004) A systems view of mRNP biology. 
Castello A, Fischer B, Eichelbaum K et al (2012) Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
